Molecular determinants of Arabidopsis quantitative disease resistance to the fungal pathogen Sclerotinia sclerotiorum
Because of their complex interplay and their small phenotypic effect, the functional characterization of QDR genes remains limited. How broad host range necrotrophic fungi manipulate plant programmed cell death is for instance largely unknown.
We designed a time-resolved automated disease phenotyping pipeline and assessed the kinetics of disease symptoms with unprecedented resolution. Comparing polymorphisms in resistant and susceptible A. thaliana accessions, we identified alleles of the nucleotide-binding site leucine-rich repeat gene LAZ5 in the resistant accessions Rubenzhnoe and Lip-0. Two LAZ5-deficient mutant lines in the Col-0 genetic background showed enhanced QDR to S. sclerotiorum (Barbacci et al. 2018). These findings illustrate the value of time-resolved image-based phenotyping for unravelling the genetic bases of complex traits such as QDR.
We designed a time-resolved automated disease phenotyping pipeline and assessed the kinetics of disease symptoms with unprecedented resolution. Comparing polymorphisms in resistant and susceptible A. thaliana accessions, we identified alleles of the nucleotide-binding site leucine-rich repeat gene LAZ5 in the resistant accessions Rubenzhnoe and Lip-0. Two LAZ5-deficient mutant lines in the Col-0 genetic background showed enhanced QDR to S. sclerotiorum (Barbacci et al. 2018). These findings illustrate the value of time-resolved image-based phenotyping for unravelling the genetic bases of complex traits such as QDR.
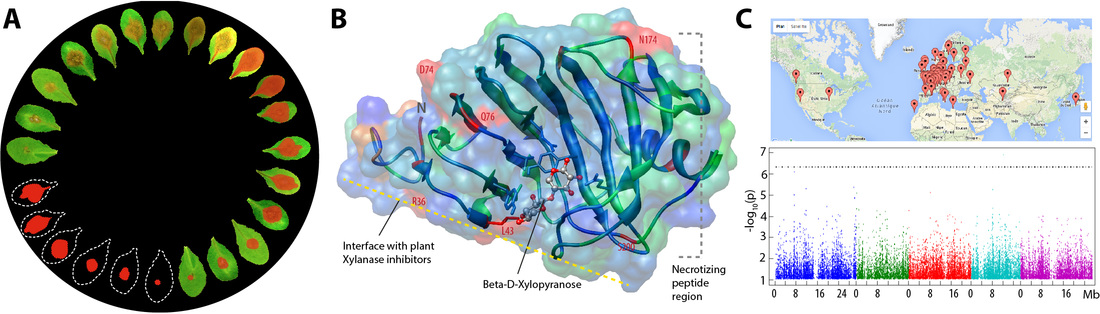
Figure legend : A) The range of symptoms on Arabidopsis thaliana leaves inoculated by S. sclerotiorum with an illustration of image processing used to quantify disease. B) Molecular structure of a fungal protein predicted to facilitate host colonization. C) Genome wide association mapping of plant genes contributing to quantitative disease resistance, with a world map indicating the origin of plant accessions used.
Selected reading:
Rapid identification of an Arabidopsis NLR gene as a candidate conferring susceptibility to Sclerotinia sclerotiorum using time-resolved automated phenotyping. Barbacci et al. Plant J. 2020
Advances on plant-pathogen interactions from molecular toward systems biology perspectives. Peyraud et al. Plant J. 2016
Resistance to phytopathogens e tutti quanti: placing plant quantitative disease resistance on the map. Roux et al. Mol Plant Pathol. 2014
Collaborations:
Contact: Sylvain Raffaele, Adelin Barbacci, Mehdi Khafif
Rapid identification of an Arabidopsis NLR gene as a candidate conferring susceptibility to Sclerotinia sclerotiorum using time-resolved automated phenotyping. Barbacci et al. Plant J. 2020
Advances on plant-pathogen interactions from molecular toward systems biology perspectives. Peyraud et al. Plant J. 2016
Resistance to phytopathogens e tutti quanti: placing plant quantitative disease resistance on the map. Roux et al. Mol Plant Pathol. 2014
Collaborations:
Contact: Sylvain Raffaele, Adelin Barbacci, Mehdi Khafif
We used GWA mapping in A. thaliana natural populations to identify genetic loci associated with QDR to S. sclerotiorum (Ss). Using insertion mutant lines, we confirmed that the POQR (Prolyl-oligopeptidase involved in Quantitative Resistance) and ARPC4 (actin-related protein complex 4) genes confer enhanced QDR to Ss. We showed that POQR function is conserved in A. thaliana (Brassicaceae) and tomato (Solanaceae) and that similar sequence and expression changes shaped the evolution of this gene in multiple plant lineages. Our study identified a previously uncharacterized gene mediating QDR against S. sclerotiorum. It showed that some QDR determinants are conserved in distantly related plants (Badet et al. 2017).
Analysis of natural diversity at the ARPC4 locus in 52 A. thaliana accessions revealed two clear haplotypes, associated with different levels of resistance to S. sclerotiorum. The resistant haplotype is characterized by a ~1Kbp insertion upstream of the ARPC4 gene, associated with an increase in ARPC4 gene expression upon Ss infection (Badet et al. 2018).
Analysis of natural diversity at the ARPC4 locus in 52 A. thaliana accessions revealed two clear haplotypes, associated with different levels of resistance to S. sclerotiorum. The resistant haplotype is characterized by a ~1Kbp insertion upstream of the ARPC4 gene, associated with an increase in ARPC4 gene expression upon Ss infection (Badet et al. 2018).
Badet T, Voisin D, Mbengue M, Barascud M, Sucher J, Sadon P, Balagué C, Roby D, Raffaele S. Parallel evolution of the POQR prolyl oligo peptidase gene conferring plant quantitative disease resistance. PLoS Genet. 2017 Dec 22;13(12):e1007143. https://journals.plos.org/plosgenetics/article?id=10.1371/journal.pgen.1007143
Relationship between quantitative immunity and signals of the environment - Thigmoimmunity : priming of immunity against Sclerotinia by mechanical cues
In natural settings, the combined action of environmental cues such as wind or rain and the colonization of host tissues by filamentous pathogens generate complex mechanical signals in plant tissues that can modify the outcome of plant-pathogen interactions. We found that plant mechano-responsive genes are induced in later stages of colonization by Ss. Whether deformations of plant tissues caused by fungal colonization induce mechanical signals interfering with QDR remains elusive.
Mechanical stimulation by touching leaves repeatedly increases Arabidopsis resistance to subsequent infections by B. cinerea, a phenomenon known as defense priming (Benikhlef et al. 2013). We verified that mechanical priming confers enhanced QDR to Ss in A. thaliana. The use of mechanical waves is adapted for tightly controlled high-throughput mechanical stimulation of plants.
Mechanical stimulation by touching leaves repeatedly increases Arabidopsis resistance to subsequent infections by B. cinerea, a phenomenon known as defense priming (Benikhlef et al. 2013). We verified that mechanical priming confers enhanced QDR to Ss in A. thaliana. The use of mechanical waves is adapted for tightly controlled high-throughput mechanical stimulation of plants.
Links:
http://journal.frontiersin.org/article/10.3389/fpls.2016.00422/full
Mechanoperception and thigmomorphogenesis at INRA
http://www6.ara.inra.fr/piaf_eng/About/Teams/MECA
Selected reading:
Mbengue M, Navaud O, Peyraud R, Barascud M, Badet T, Vincent R, Barbacci A, Raffaele S. Emerging Trends in Molecular Interactions between Plants and the Broad Host Range Fungal Pathogens Botrytis cinerea and Sclerotinia sclerotiorum. Front Plant Sci. 2016 Mar 31;7:422.
http://journal.frontiersin.org/article/10.3389/fpls.2016.00422/full
Barbacci A, Magnenet V, Lahaye M. Thermodynamical journey in plant biology. Front Plant Sci. 2015 Jun 30;6:481.
http://journal.frontiersin.org/article/10.3389/fpls.2015.00481/full
Contact: Adelin Barbacci
http://journal.frontiersin.org/article/10.3389/fpls.2016.00422/full
Mechanoperception and thigmomorphogenesis at INRA
http://www6.ara.inra.fr/piaf_eng/About/Teams/MECA
Selected reading:
Mbengue M, Navaud O, Peyraud R, Barascud M, Badet T, Vincent R, Barbacci A, Raffaele S. Emerging Trends in Molecular Interactions between Plants and the Broad Host Range Fungal Pathogens Botrytis cinerea and Sclerotinia sclerotiorum. Front Plant Sci. 2016 Mar 31;7:422.
http://journal.frontiersin.org/article/10.3389/fpls.2016.00422/full
Barbacci A, Magnenet V, Lahaye M. Thermodynamical journey in plant biology. Front Plant Sci. 2015 Jun 30;6:481.
http://journal.frontiersin.org/article/10.3389/fpls.2015.00481/full
Contact: Adelin Barbacci
Molecular mechanisms and evolution of broad host range pathogenicity in fungi
Fungal plant pathogens produce secreted proteins adapted to function outside fungal cells to facilitate colonization of their hosts. For fungi from the Sclerotiniaceae family the repertoire and function of secreted proteins remains elusive (Mbengue et al., 2016). We combined bioinformatics approaches to determine the repertoire of Ss effector candidates and conducted detailed sequence and expression analyses on selected candidates. We identified 486 Ss secreted protein genes expressed sunflower. Among those, we identified 78 effector candidates based on canonical effector protein properties (Guyon et al., 2014). We analyzed amino acids usage and intrinsic protein disorder in three Sclerotiniaceae species to reveal signatures associated with effector candidates (Badet et al. 2015). A total of 22 effector candidates have been cloned and selected for functional analysis. To facilitate genomic analyses of virulence determinants in Ss, we generated a complete genome sequence of Ss from Single Molecule Real-Time Sequencing technology with highly accurate annotations produced using an extensive RNA sequencing data set (Derbyshire et al. 2017). Using this high-quality assembly as a reference, we re-sequenced the genomes of 25 isolates of Ss collected from four different continents and conducted SNP-based analyses of population structure and selective sweeps (Derbyshire et al. 2019).
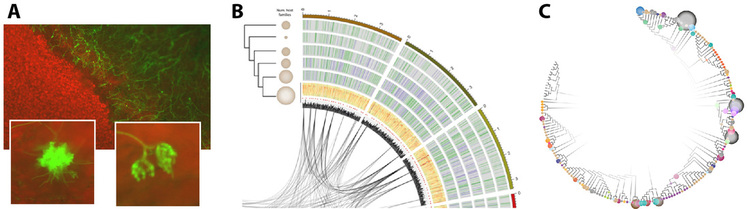
Figure legend : A) S. sclerotiorum colonizing an Arabidopsis leaf imaged using a strain expressing the green fluorescent protein. B) A Circos diagram illustrating comparative genomics of fungal species related to S. sclerotiorum. C) A phylogenetic tree of fungi from the Sclerotiniaceae family.
Selected reading:
Badet T, Peyraud R, Mbengue M, Navaud O, Derbyshire M, Oliver RP, Barbacci A, Raffaele S. Codon optimization underpins generalist parasitism in fungi. Elife. 2017 Feb 3;6 https://elifesciences.org/content/6/e22472
Badet T, Peyraud R, Raffaele S. Common protein sequence signatures associate with Sclerotinia borealis lifestyle and secretion in fungal pathogens of the Sclerotiniaceae. Front Plant Sci. 2015 Sep 24;6:776.
Guyon K, Balagué C, Roby D, Raffaele S. Secretome analysis reveals effector candidates associated with broad host range necrotrophy in the fungal plant pathogen Sclerotinia sclerotiorum. BMC Genomics. 2014 May 4;15:336.
Contact : Sylvain Raffaele
Badet T, Peyraud R, Mbengue M, Navaud O, Derbyshire M, Oliver RP, Barbacci A, Raffaele S. Codon optimization underpins generalist parasitism in fungi. Elife. 2017 Feb 3;6 https://elifesciences.org/content/6/e22472
Badet T, Peyraud R, Raffaele S. Common protein sequence signatures associate with Sclerotinia borealis lifestyle and secretion in fungal pathogens of the Sclerotiniaceae. Front Plant Sci. 2015 Sep 24;6:776.
Guyon K, Balagué C, Roby D, Raffaele S. Secretome analysis reveals effector candidates associated with broad host range necrotrophy in the fungal plant pathogen Sclerotinia sclerotiorum. BMC Genomics. 2014 May 4;15:336.
Contact : Sylvain Raffaele