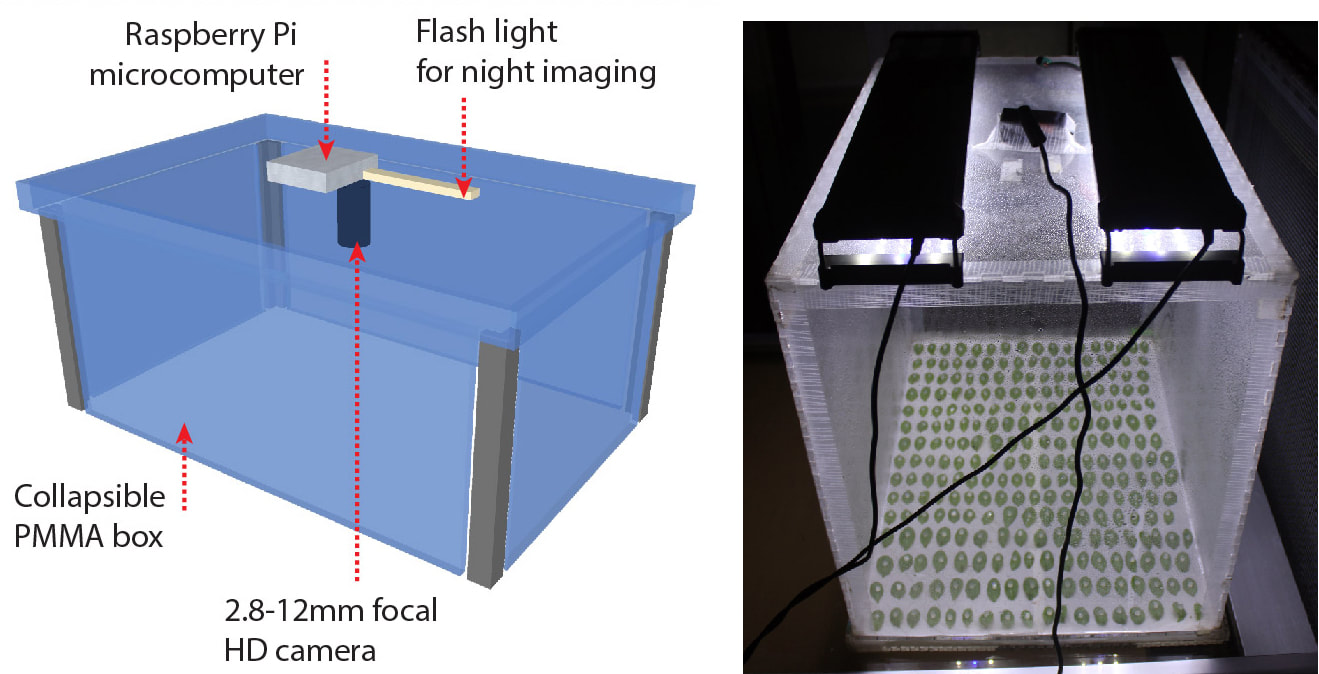
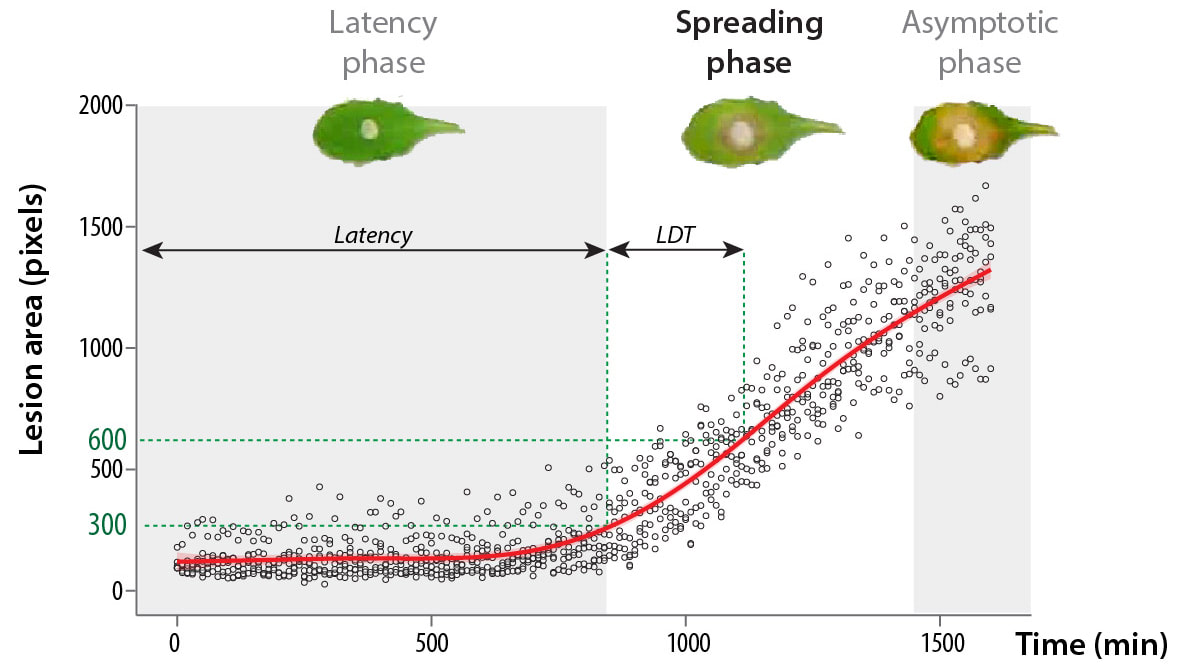
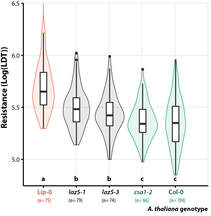
| You may miss the lab in these days of lockdown. But have you ever wished you could spend the week end with something else than checking symptoms in the lab ? Ever thought "sh*t I should have pictured my plants two hours ago"? We designed a simple and affordable solution for versatile plant disease phentoyping over time. All you need is a plexiglass box, a cheap camera and raspberryPi. We provide guidance for the design, and the code needed to make it a convenient phenotyping machine! Extracting meaningful information from leaf images in a high throughput can be daunting. This simple setup allows capturing and analysing disease lesions automatically over time, giving access to plant disease kinetics parameters. We designed the first prototype in 2014 and kept improving it since then. The possibilities for improvement are endless, your suggestions are welcome! Thanks to this tool, we found that lesion doubling time is a key parameter to describe plant resistance to the fungal pathogen Sclerotinia. The high resolution offered by high-throughput image analysis, we reveal that the Arabidopsis NLR gene LAZ5, but not the closely related CSA1, confers susceptibility to Sclerotinia. You will find a tutorial for this tool here:https://github.com/A02l01/INFEST with the code (& more tuto and test files) here:https://onlinelibrary.wiley.com/doi/abs/10.1111/tpj.14747 [7/7] |
|
0 Comments
|
Archives
November 2023
Categories |

 RSS Feed
RSS Feed
