We report a comparative analysis of natural selection for codon optimization across fungi, revealing that it associates with the range of hosts parasites can infect. The analysis includes parasites that are major threats to human and animal health and to food security worldwide. The full report is available at:
https://elifesciences.org/content/6/e22472
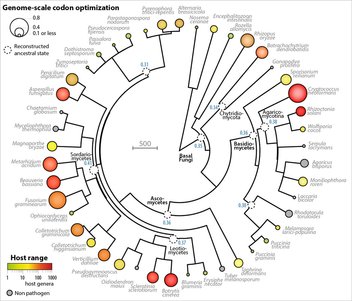
| From the abstract: We show that codon optimization underlies genome adaptation in broad host range parasite. The longer proteins encoded by broad host range fungi likely increase natural selection on codon optimization in these species. Accordingly, codon optimization correlates with host range across the fungal kingdom (figure on the left). At the species level, biased patterns of synonymous substitutions underpin increased codon optimization in a generalist but not a specialist fungal pathogen. Virulence genes were consistently enriched in highly codon-optimized genes of generalist but not specialist species. We conclude that codon optimization is related to the capacity of parasites to colonize multiple hosts. Our results link genome evolution and translational regulation to the long term persistence of generalist parasitism. Image: Patterns of codon substitutions in a generalist fungal pathogen (vertical, greenish colors) and in a specialist fungal pathogen (horizontal, blueish colors). INRA press release: http://www.toulouse.inra.fr/Toutes-les-actualites/L-incroyable-capacite-adaptative-des-champignons-pathogenes-pour-infecter-de-multiples-especes |

 RSS Feed
RSS Feed
