The fungal pathogen Sclerotinia sclerotiorum infects hundreds of plant species, including members of the Brassicaceae and Solanaceae families. These two botanical families diverged over 100 million years ago, whereas Sclerotinia lineage gained ability to infect them less than 10 million years ago. Is there a chance that today's Brassicacea and Solanaceae developped similar defense responses to S. sclerotiorum in parallel?
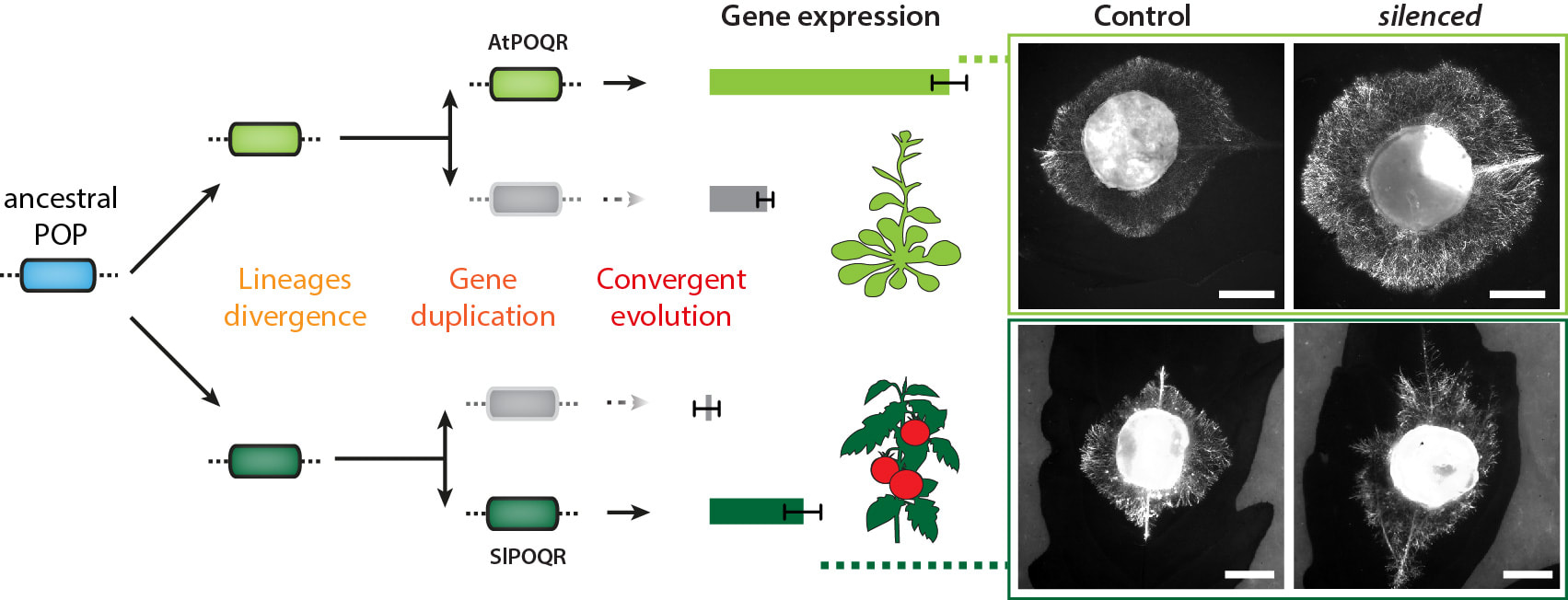
Yes! we found one example. But we had to identify genes controlling quantitative disease resistance to S. sclerotiorum in Arabidopsis (Brassicaceae) first. Using genome wide association mapping and reverse genetics, we found a prolyl-oligopeptidase (an enzyme predicted to cleave small peptides after a proline) that is important for disease resistance. We named it POQR, and played it as a wild card. Loss of this gene compromised QDR against S. sclerotiorum but not against a bacterial pathogen. Natural diversity analysis associated POQR sequence with QDR. Remarkably, the same amino acid changes occurred after independent duplications of POQR in ancestors of multiple plant species, including A. thaliana and tomato. Comparative analysis of global transcriptome data from S. sclerotiorum-infected A. thaliana and tomato plants showed that POQR evolved induction upon infection after gene duplication independently in the two species, and that divergence of gene expression following duplication is widespread at the genome scale.
Read the full story here: http://journals.plos.org/plosgenetics/article?id=10.1371/journal.pgen.1007143

 RSS Feed
RSS Feed
