academic.oup.com/femsre/advance-article/doi/10.1093/femsre/fuab002/6095737
The original idea and a lot of the hard work came from my amazing chinese co-authors. I was delighted to work again with my fellow LFCat Suomeng!
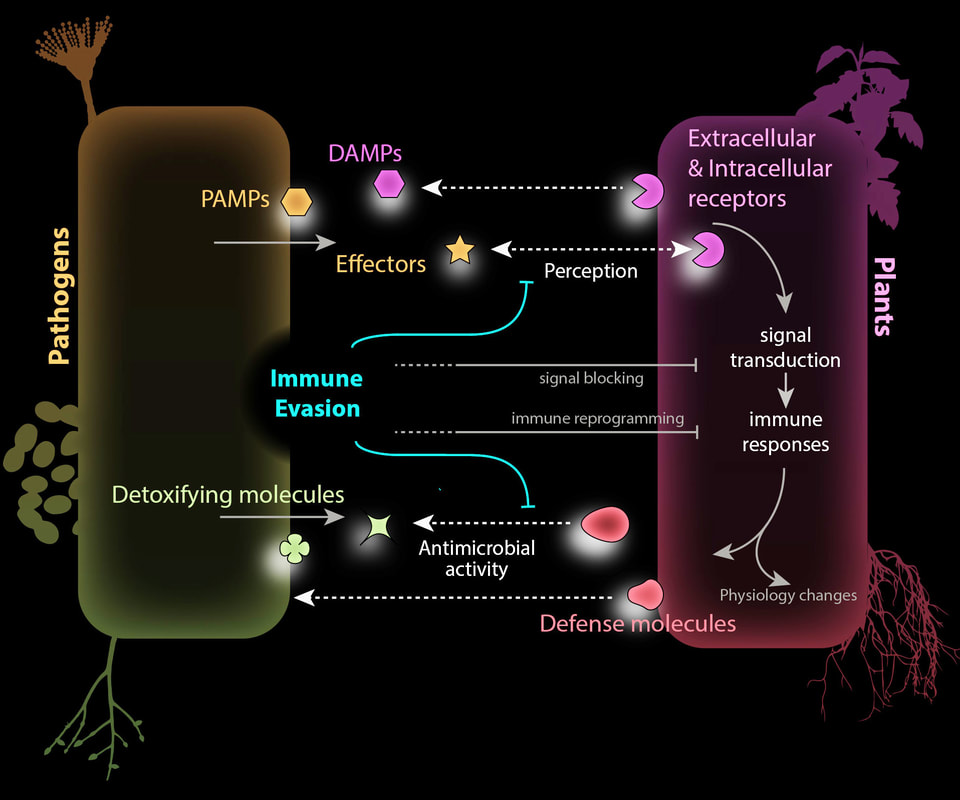
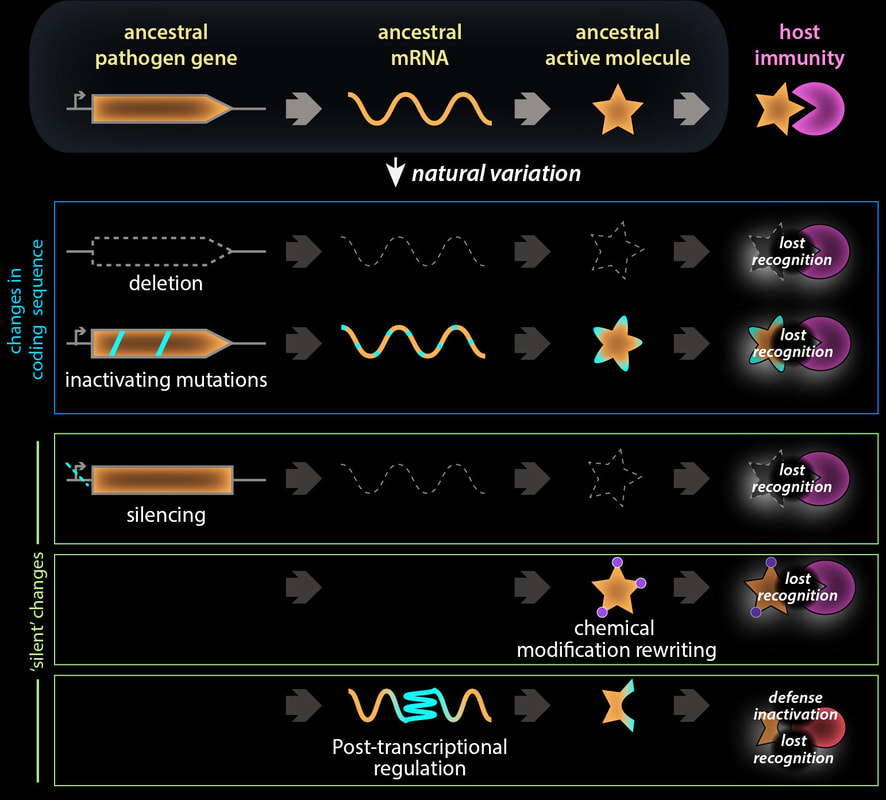
Traditionally, it is believed that pathogens mainly rely on sequence mutations to obtain new capability to evade host immune recognition. Indeed, the development of high-throughput sequencing technology provided great help to screen and revealed mutations at the nucleotide sequence level. On the other hand, the recent discovery of natural variations beyond genetic sequence changes uncovered additional layers by which plant pathogens escape host immune selective pressure. In this paper, we discuss biochemical modifications of PAMPs, post-transcriptional regulation of virulence, and gene silencing of effectors that allow pathogens to evade plant immune systems. We still lack robust and cost-effective profiling methods to detect these changes at the population level. Besides the application of novel technologies, novel analytical approaches are needed.

 RSS Feed
RSS Feed
