Read the full story here: http://onlinelibrary.wiley.com/doi/10.1111/mec.14523/full/
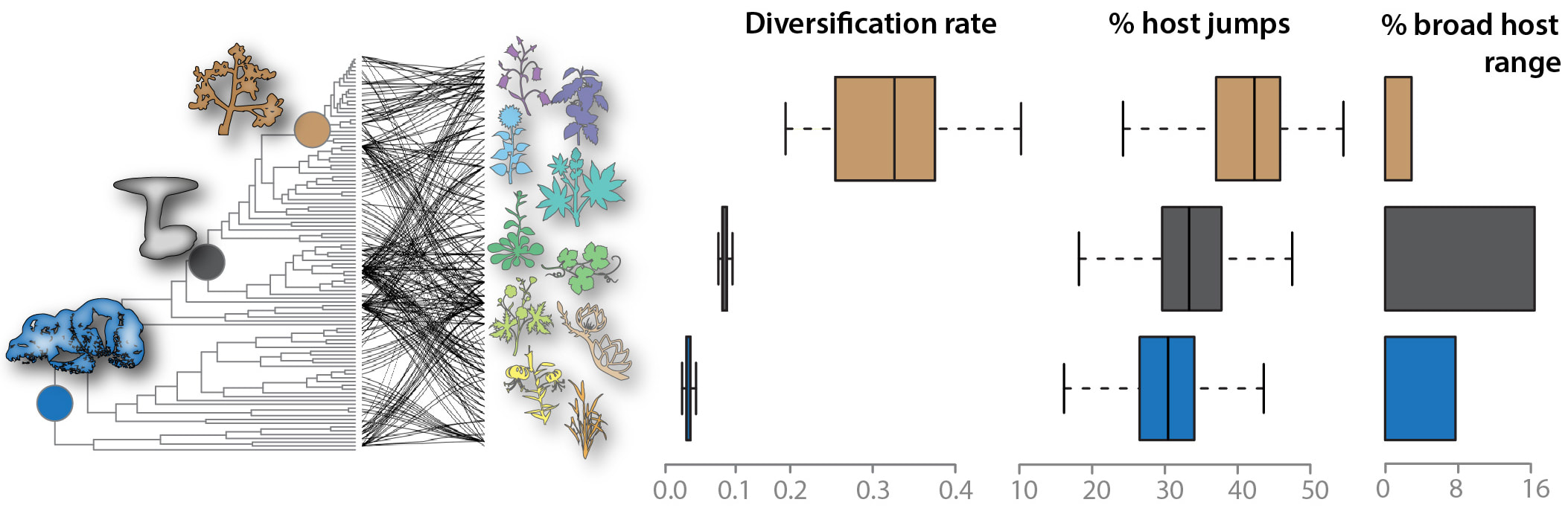
Shifts in diversification rates and host jump frequencies in Sclerotiniaceae fungal plant pathogens2/8/2018 The fungal pathogen Sclerotinia sclerotiorum is notorious for its ability to infect hundreds of plant species, but its close relatives do not share this property. Studying how the "broad host range" trait evolved in the Sclerotiniaceae family would pave the way for resolving the underlying molecular bases. Using a phylogenetic framework, we associate diversification rates, the frequency of host jump events, and host range variation during the evolution of this family. Variations in diversification rate during the evolution of the Sclerotiniaceae define three major macro-evolutionary regimes with contrasted proportions of species infecting a broad range of hosts. Host-parasite co-phylogenetic analyses pointed towards parasite radiation on distant hosts long after host speciation (host jump or duplication events) as the dominant mode of association with plants in the Sclerotiniaceae. The intermediate macro-evolutionary regime showed a low diversification rate, high frequency of duplication events, and the highest proportion of broad host range species.
Read the full story here: http://onlinelibrary.wiley.com/doi/10.1111/mec.14523/full/
0 Comments
Our latest study appears today in Plos Genetics!
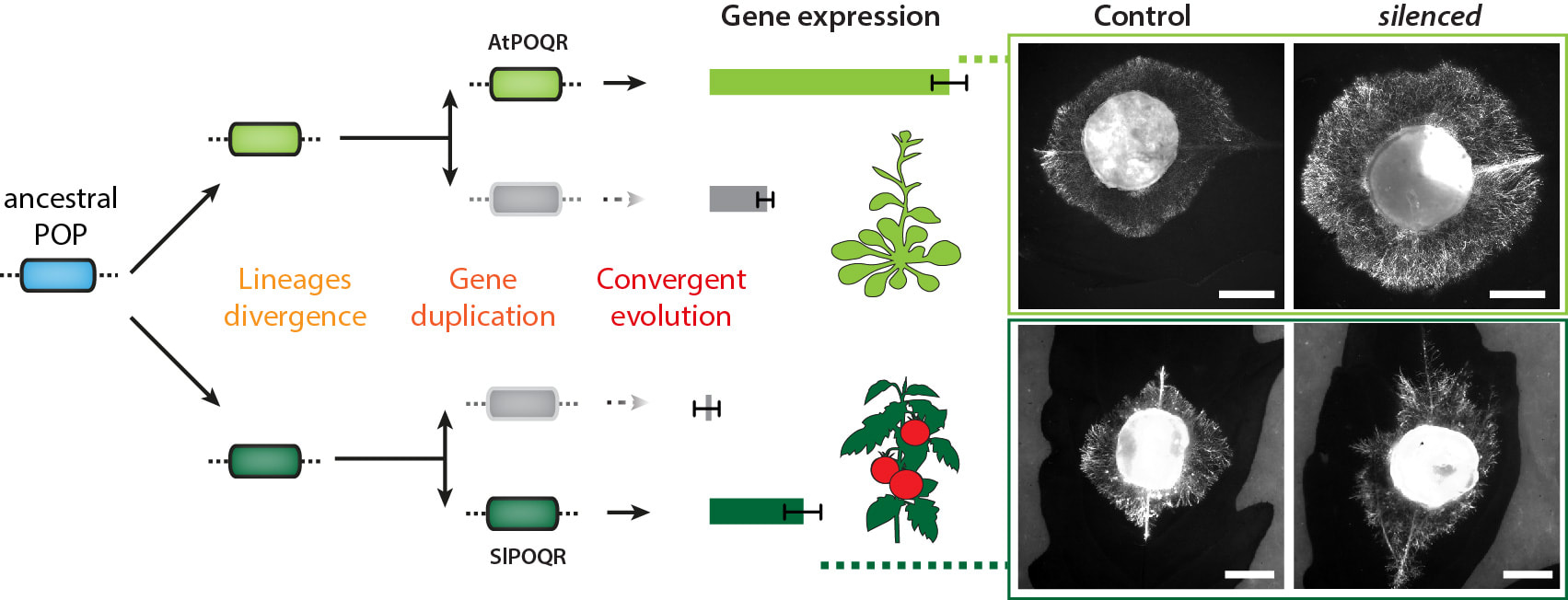
The fungal pathogen Sclerotinia sclerotiorum infects hundreds of plant species, including members of the Brassicaceae and Solanaceae families. These two botanical families diverged over 100 million years ago, whereas Sclerotinia lineage gained ability to infect them less than 10 million years ago. Is there a chance that today's Brassicacea and Solanaceae developped similar defense responses to S. sclerotiorum in parallel? Yes! we found one example. But we had to identify genes controlling quantitative disease resistance to S. sclerotiorum in Arabidopsis (Brassicaceae) first. Using genome wide association mapping and reverse genetics, we found a prolyl-oligopeptidase (an enzyme predicted to cleave small peptides after a proline) that is important for disease resistance. We named it POQR, and played it as a wild card. Loss of this gene compromised QDR against S. sclerotiorum but not against a bacterial pathogen. Natural diversity analysis associated POQR sequence with QDR. Remarkably, the same amino acid changes occurred after independent duplications of POQR in ancestors of multiple plant species, including A. thaliana and tomato. Comparative analysis of global transcriptome data from S. sclerotiorum-infected A. thaliana and tomato plants showed that POQR evolved induction upon infection after gene duplication independently in the two species, and that divergence of gene expression following duplication is widespread at the genome scale. Read the full story here: http://journals.plos.org/plosgenetics/article?id=10.1371/journal.pgen.1007143 Thomas succesfully defended his PhD thesis!! congrats Dr. Badet!
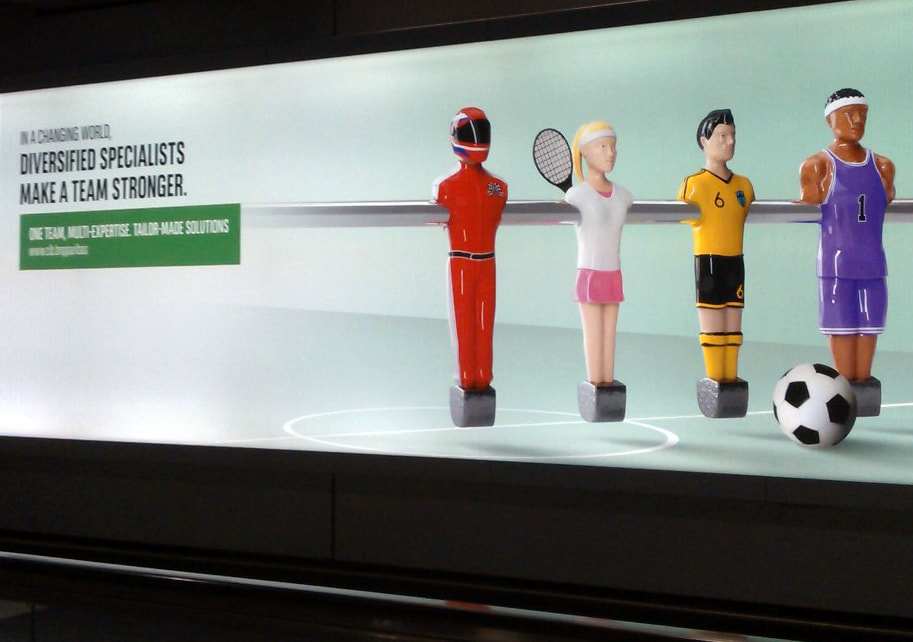
Many thanks to Sabine Fillinger (INRA BIOGER), Mathilde Fagard (INRA Versailles), Valérie Geffroy (Institute of Plant Sciences Paris-Saclay), Daniel Croll (Université Neuchatel), Jean Benoit Morel (INRA BGPI Montpellier) and Christophe Roux (LRSV Toulouse) for your insights and enthusiatic discussions. On the next day: fascinating seminar by Prof. Daniel Croll on "How pathogens rapidly surmount plant resistance in agricultural ecosystems". Very interesting points on the evolution of fungicide resistance, the power of GWAS in fungal pathogens and revelations from Zymoseptoria pangenome. Check Daniel's website for more: http://www.pathogen-genomics.org/ Amazing full day visit @Max Planck Institute for Plant Breeding Research in Cologne, with very inspiring discussions before and after my seminar. Many thanks to Stephane Hacquard for the invitation, Thomas Nakano, Bhandari Deepak and Joram Dongus, Hirofumi Nakagami, Kenichi Tsuda, Takaki Meakawa, Miltos Tsiantis, Paul Schulze-Lefert and the PhD students for sharing views and fantastic science. The day was so packed that I nearly missed the cathedral on my way to the train station... Spotted in Frankfurt airport while commuting: an add supporting the "polyspecialist" view of broad host range pathogens ("diversified specialist genes make a pathogen stronger") ?...
One day in Bordeaux with Sebastien Mongrand to attend lab seminar by Sophien Kamoun. The first opportunity to catch up with my two post-doc mentors at the same time!
check the LBM (membrane biogenesis lab @INRA Bordeaux): http://www.biomemb.cnrs.fr/
Joint annual meeting of the "Resistance" and "Effectome" INRA networks in Banyuls, FR. With a talk by Remi and Sylvain @QIPlab.
Great science and a fantastic travel experience for the 2nd Lazy Fat Cats meeting in Hohhot, inner Mongolia, hosted by Ruofang Zhang and Sumeng Dong.
The program included talks by Sophien Kamoun and Chih-hang Wu (TSL Norwich), Yuanchao Wang and Suomeng Dong (Nanjing Agricultural University), Edgar Huitema (James Hutton Institute), Wenbo Ma (UC Riverside) , Zhenyu Liu (Anhui Agricultural University), Kentaro Yoshida (Kobe University), Jian-min Zhou (Chinese Academy of Science), Rosa Lozano-Duran and Alberto Macho (Shangaï University), Sebastian Schornack (TSL Cambridge). Many thanks to our hosts for this amazing trip, and to the old and new LFCs for keeping the spirit alive! Looking forward to LFC#3!
Our project FUSSPOT "Impact of fungal secreted signalling peptides on plant symbioses" receives funding by the federative institute of research (FR AIB) and TULIP. This is a collaborative project initiated by Nicolas Frei-dit-Frey @ LRSV (https://www.lrsv.ups-tlse.fr/?-Themes-de-recherche,80-) aiming at exploring the biological activity of fungal small secreted peptides on plant interactions with Rhizophagus irregularis and Sclerotinia sclerotiorum.
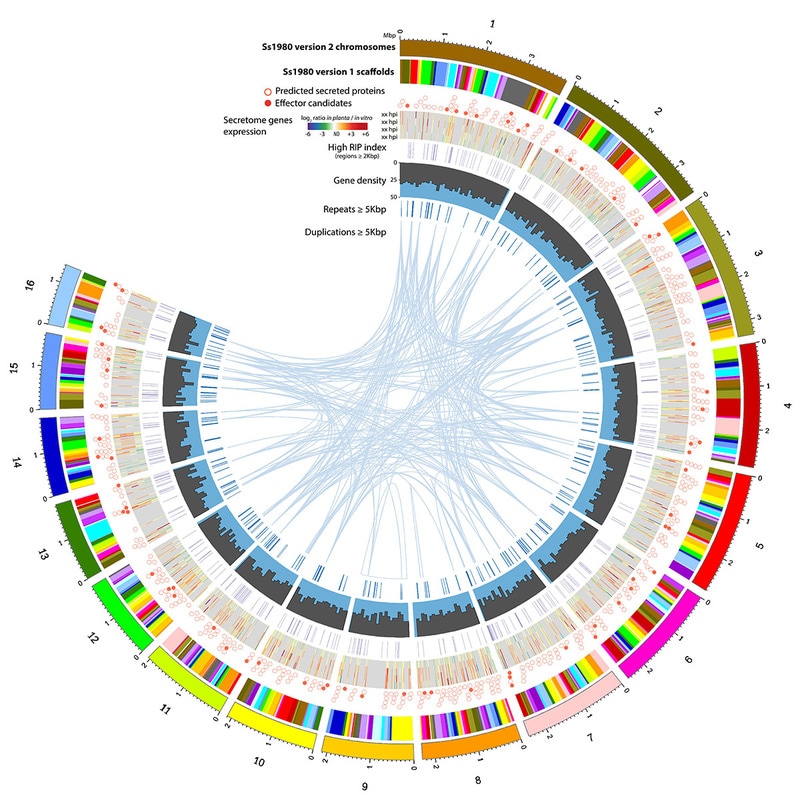
As a part of an international consortium led by the group of Richard Oliver from Uni. Curtin (Austalia), we contributed to the assembly and analysis of the complete genome sequence of our favourite fungus Sclerotinia sclerotiorum. The article is published today in Genome Biology and Evolution: https://academic.oup.com/gbe/article-lookup/doi/10.1093/gbe/evx030
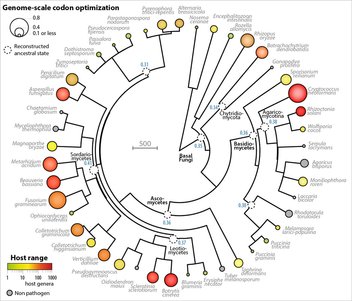
Sclerotinia is featured in eLife, with a bunch of other fungi! We report a comparative analysis of natural selection for codon optimization across fungi, revealing that it associates with the range of hosts parasites can infect. The analysis includes parasites that are major threats to human and animal health and to food security worldwide. The full report is available at: https://elifesciences.org/content/6/e22472
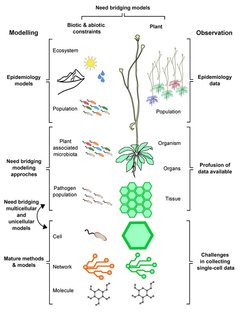
Our review on the use of systems biology to study plant-pathogen interactions is online: Peyraud R, Dubiella U, Barbacci A, Genin S, Raffaele S, Roby D. Advances on plant-pathogen interactions from molecular toward systems biology perspectives. Plant J. 2016 Nov 21.
20/01/17: The post-doc positions are filled. Thanks for applying Job offer update! We now have 2 Post-doc and 1 research assistant positions available and fully funded Post doc positions are available immediately for up to 2 years (until november 2018), the RA position will be available from June 2017. A major objective in our lab is to decipher the genetic architecture of plant quantitative immunity. We are especially focusing on the diversity of quantitative immunity program in multiple plant genotypes and species, and on the connections between immunity and the response to abiotic stress. We also develop a research program aiming at understanding how the fungal pathogen S. sclerotiorum manipulates plant cells to colonize host tissues, and how its virulence program evolves in the context of plant quantitative immunity. Our approaches are highly multidisciplinary involving methods ranging from molecular and cell biology, genetics, high throughput phenotyping of plants and microbes, genomics and transcriptomics, systems biology and modeling. We are looking for highly motivated and talented collaborators with expertise in these approaches. Experience in plant pathology or fungal microbiology will be seen as an advantage. For further information, please contact project leader Dr. Sylvain Raffaele +33 561 285 326, [email protected] Apply by contacting Sylvain Raffaele directly or via the LIPM application system (https://www6.toulouse.inra.fr/lipm/Opportunites-Formations/Candidater).
 Thomas and Sylvain are spending 3 days at the Auberge du Cèdre near Montpellier for the 3rd meeting of the "Resistance Network". This is a national meeting sponsored by INRA on all aspects of plant resistance, with a special emphasis this year on durability and biocontrol. Thomas is presenting his PhD work on the evolution of a tradeoff between quantitative immunity and abiotic stress response.
|
Archives
November 2023
Categories |
|||||||||||||||||||||||||||||||












 RSS Feed
RSS Feed
